Library synthesis
Batch retrosynthesis for analog series and arbitrary molecule lists.
Libraries are batches of molecules submitted for retrosynthesis as a group. Each entry gets its own task; convergence analysis runs automatically across the result set. Manage libraries from the Library sidebar entry.
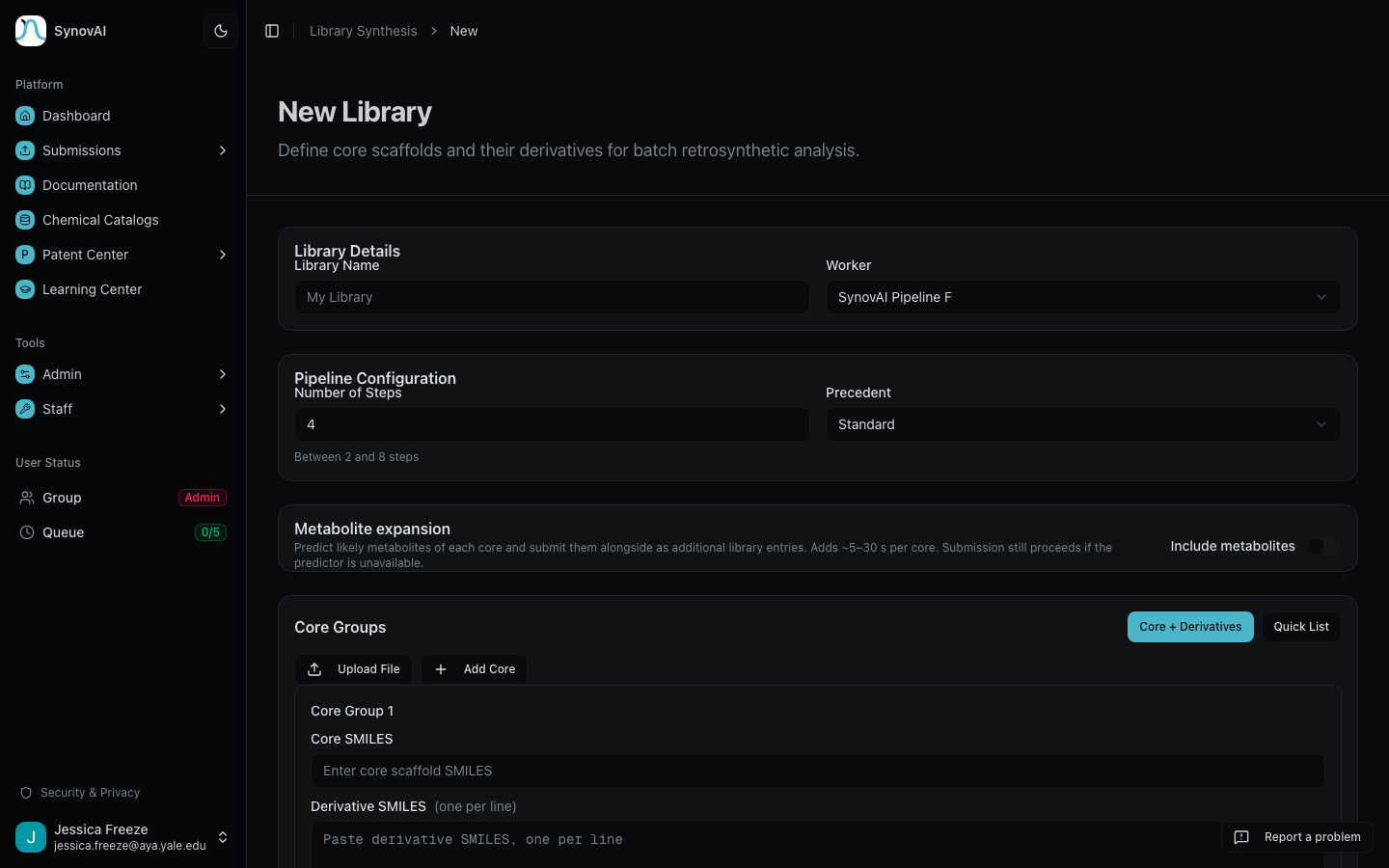
Two ways to build one
Core + Derivatives
Define a core scaffold and specify derivative positions. Best for systematic SAR exploration around a common intermediate.
Quick List
Paste a flat list of SMILES (one per line) and each line becomes its own library entry. Best for batch retrosynthesis of related or unrelated molecules, or for piping whitespace selections from Patent Fence.
The DRAFT lifecycle
Every new library is created in DRAFT status. While the library is in DRAFT you can:
- Rename the library
- Add or remove member molecules
- Reorder members
- Change synthesis parameters (step count, precedent level, starting material, commercial stock settings)
These four operations are gated server-side on status === 'DRAFT'.
The moment you click Submit All, the library transitions through
SUBMITTING into RUNNING, and the edit endpoints reject further writes.
Why edits lock at submission
Once a library is submitted, every member gets its own Pipeline F job queued at the parameters captured at submission time. Allowing edits mid-run would produce a result set where some entries reflect the new parameters and some reflect the old ones, with no way to tell which is which from the dashboard. Locking on submission keeps the result set internally consistent and the audit log honest.
If you need to change a parameter after submission, clone the library (any of its results stays put) and edit the clone in DRAFT.
Submit all
From DRAFT, click Submit All to launch retrosynthesis on every member. Track progress on the library dashboard. Libraries auto- complete when all member tasks finish, and convergence analysis runs automatically on the result set so you don't have to launch it manually.
Shared intermediates
Use convergence analysis across library results to find shared intermediates, the key building blocks that appear across multiple analog routes. These enable more efficient parallel synthesis strategies, where you make the shared intermediate once and diverge to the analogs.
Manage libraries
Each library card on the dashboard is clickable to open the result set. Delete a library with the trash button (with confirmation); the underlying tasks are unaffected unless you choose to archive them separately.
What to read next
- Medicinal chemist reading path: SAR-focused workflows that lean heavily on libraries.
- Convergence explorer: navigating shared intermediates in detail.
- Patent tools: where Patent Fence whitespace selections turn into a library submission.